 |
Influenza Virus Binds to Its Host Cell Through Multiple Dynamic Interactions
Professor Andreas Herrmann
Humboldt University of Berlin, Department of Biology, Berlin, Germany
3:30 pm Tuesday, 16 September 2014
EN313 Lecture Theatre (EN Building), Hawthorn.
Abstract
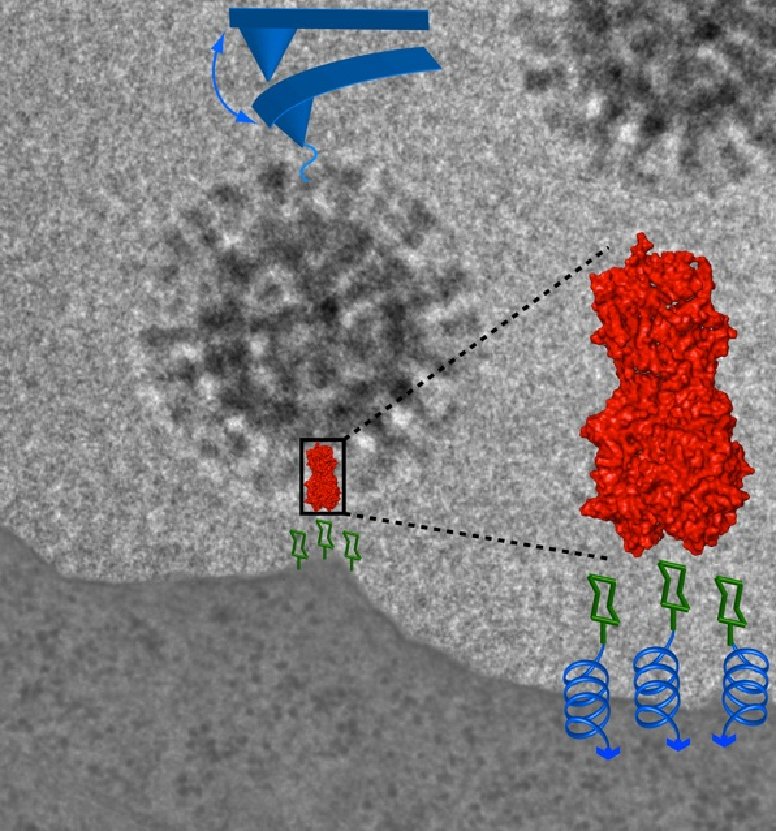 Influenza virus belongs to a wide range of enveloped viruses. Virus-host cell binding marks the first critical step of infection. Hence, forces involved in this process are essential. The major spike protein hemagglutinin binds sialic acid residues of glycoproteins and lipids with dissociation constants in the millimolar range (Sauter et al. (1992) Biochemistry 31:9609–9621), indicating a multivalent binding mode. We characterized the attachment of influenza virus to host cell receptors using three independent approaches. Optical tweezers and atomic force microscopy-based single molecule force spectroscopy provides powerful tools to measure binding forces in biological systems. Optical tweezers and AFM-based single molecule force spectroscopy revealed very low interaction forces. The observation of sequential unbinding events strongly suggests a multivalent binding mode between virus and cell membrane. However, an assignment of forces to their underlying molecular interactions involved in these processes is difficult or even cannot be obtained by these techniques. In molecular dynamics, time-dependent interactions between all atoms within a given system are calculated numerically within timescales smaller than those observed in biological phenomena. Force probe molecular dynamics simulations provide a way to overcome this time limitation by introducing a moving harmonic potential as a "virtual spring" acting on given atoms. Molecular dynamics simulations reveal a variety of unbinding pathways that indicate a highly dynamic interaction between HA and its receptor allowing to rationalize the binding of influenza virus to host cells quantitatively at molecular level. Influenza virus belongs to a wide range of enveloped viruses. Virus-host cell binding marks the first critical step of infection. Hence, forces involved in this process are essential. The major spike protein hemagglutinin binds sialic acid residues of glycoproteins and lipids with dissociation constants in the millimolar range (Sauter et al. (1992) Biochemistry 31:9609–9621), indicating a multivalent binding mode. We characterized the attachment of influenza virus to host cell receptors using three independent approaches. Optical tweezers and atomic force microscopy-based single molecule force spectroscopy provides powerful tools to measure binding forces in biological systems. Optical tweezers and AFM-based single molecule force spectroscopy revealed very low interaction forces. The observation of sequential unbinding events strongly suggests a multivalent binding mode between virus and cell membrane. However, an assignment of forces to their underlying molecular interactions involved in these processes is difficult or even cannot be obtained by these techniques. In molecular dynamics, time-dependent interactions between all atoms within a given system are calculated numerically within timescales smaller than those observed in biological phenomena. Force probe molecular dynamics simulations provide a way to overcome this time limitation by introducing a moving harmonic potential as a "virtual spring" acting on given atoms. Molecular dynamics simulations reveal a variety of unbinding pathways that indicate a highly dynamic interaction between HA and its receptor allowing to rationalize the binding of influenza virus to host cells quantitatively at molecular level.
Graphic by Dr. Christian Sieben
Back to 2014 programme

|